ML Brain
Tumor Detection
06 July 2025
Description
The Context: As artificial intelligence is becoming more prominent in healthcare, particularly for early diagnosis and disease detection, this project focuses on brain-related diseases such as tumors, gliomas and meningiomas. The detection of such diseases present significant diagnostic challenges because their imaging characteristics often overlap or exhibit only subtle differences.
Currently, the manual analysis of brain MRI scans is considered time-consuming and prone to human error, particularly in healthcare systems with limited resources. This project is motivated by the potential for machine learning (ML) and deep learning (DL) to provide automated, scalable and precise image classification.
All of which are to support radiologists, improve diagnostic accuracy and accelerate the time to treatment.
The Goal: The primary goal of this project is to address the challenges of brain MRI analysis by evaluating and comparing the effectiveness of four different machine learning approaches: Support Vector Machines (SVM), Convolutional Neural Networks (CNN), Convolutional Autoencoders (CAE) and Vision Transformers (ViTs).
Key Contributions
- Data Preparation & Pre-Processing
- Developed ML - Support Vector Machine (SVM)
Tech Stack
- Language: Python
- Tools & Packages: Google Colab, sklearn, matplotlib, pytesseract, opencv-python
Description - Key Features
1) Four distinct models are compared & evaluated: Support Vector Machines (SVM), Convolutional Neural Networks (CNN), Convolutional Autoencoders (CAE), and Vision Transformers (ViT)
2) Advanced Pre-processing Pipeline: The project features a robust cleaning stage that uses cryptographic hashing for automated deduplication and a specialized artifact removal process. This process uses Tesseract-based text detection and color-based masking to remove non-anatomical elements like measuring grids, text and boxes from training dataset images.
3) Specialized Medical Data Augmentation: To address the limited dataset, the system implements techniques such as elastic deformation (to mimic natural tissue variations), Gaussian and salt-and-pepper noise, gamma correction and zooming
4) Automated Hyperparameter Optimization: Each model utilizes systematic tuning methods, such as GridSearchCV for SVM, Optuna for the Vision Transformer and grid search for the CNN architecture
Description - Key Functionalities
1) Automated Image Classification: The core functionality is the ability to take raw or pre-processed MRI images and output a diagnostic label with a corresponding confidence score.
2) Feature Extraction and Representation:
- The SVM uses Histogram of Oriented Gradients (HOG) to capture edge and shape information.
- CNNs and ViTs provide automatic feature extraction, learning hierarchical spatial patterns and global context directly from raw pixels.
- The CAE learns compact latent representations of MRI slices through unsupervised learning.
3) Model Interpretability and Visualization:
- Vision Transformers provide attention maps to visualize which regions of the brain the model focused on during prediction.
- CAEs offer visual reconstructions, allowing clinicians to compare the original image with what the network has learned to verify anatomical fidelity.
4) Dynamic Training Control: The system uses early stopping and learning rate reduction on plateau to prevent overfitting and ensure the models converge at their optimal generalization point.
5) Comprehensive Evaluation Tools: Functionality is included to generate detailed classification reports (precision, recall, F1-score) and confusion matrices to identify specific class-wise biases or misclassifications
Technical Challenges & Solutions
Technical Challenges & Solutions - Limited Dataset
Issue
The project was restricted by a limited dataset size where, each of the four diagnostic categories originally contained only about 120 images. This posed a significant risk of models memorizing noise and overfitting rather than learning generalizable features.
Solution
To address this, we implemented an extensive suite of data augmentation techniques to artificially expand the training set and improve its diversity.
- Elastic deformation was applied to mimic natural variations in brain shape and structure without distorting the image into unrecognizability.
- Gaussian noise and salt and pepper noise to simulate real-world scanning imperfections and sensor problems.
- Horizontal flips and a 1.2 zoom factor were utilized to ensure the models remained robust to different patient positions and scale-invariant regarding tumor size.
- Gamma correction was employed to simulate variations in brightness and contrast and enhance the models’ sensitivity to subtle lesions that might be hidden in underexposed regions.
The implementation of these strategies resulted in a final expanded dataset of 2317 total images.
Lesson
A major lesson learnt was that rigorous data augmentation is essential for building robust and generalizable models when working with limited medical imaging data. This process allowed the models to become more invariant to noise and transformations and ultimately achieved higher classification accuracy on unseen test data.
Technical Challenges & Solutions - Data Leakage from Data Augmentation
Issue
We observed a suspicious pattern where the validation loss consistently outperformed the training loss. This indicated that the model was not genuinely generalizing but was instead benefiting from seeing variations of the same images in both the training and validation sets. These variations of the same image were generated from the Data Augmentation solution before the splitting of the dataset into training and validation datasets occured, thus causing this issue.
Solution
To fix this we developed a custom dataset splitting function. This specific workflow ensured that the data was split into separate training and validation groups before any augmentation was applied. By isolating the raw images first any augmented versions of a single image would remain strictly within the training set and not cross over into the validation data.
Lesson
The primary lesson learnt was that splitting the dataset before performing augmentation is a fundamental requirement for maintaining the integrity of the machine learning process. This approach prevents artificial inflation of performance metrics and ensures that the validation set remains a true test of the model’s ability to handle unseen data. This experience provided a deeper understanding of how each component of a model pipeline contributes to its final reliability and performance.
Technical Challenges & Solutions - Cleaning Dataset Artifacts
Issue
Raw MRI scans contained various non-anatomical artifacts including white lines, borders, measuring grids and text overlays that could introduce bias or non-relevant features during model training.
These artifacts were likely introduced by imaging equipment or during post-processing and do not correspond to any actual anatomical features. During manual inspection, we found these overlays appeared primarily at the margins of the images in classes like pituitary, notumor and meningioma. The core challenge was that such features could introduce unintended bias or cause the models to focus on non-relevant markers instead of the actual brain tissue.
Solution
To resolve this issue we developed an automated cleaning pipeline to identify and mask these artifacts:
1) The process began by converting images to grayscale and HSV color spaces to help identify bright border pixels and colored markers using thresholding and morphological operations.
2) To protect the integrity of the diagnostic data a central brain exclusion zone was defined to ensure that no masking occurred within the actual brain tissue.
3) The pipeline also integrated Tesseract-based text detection to find and remove textual annotations and used direct edge sweeping to catch remaining border pixels.
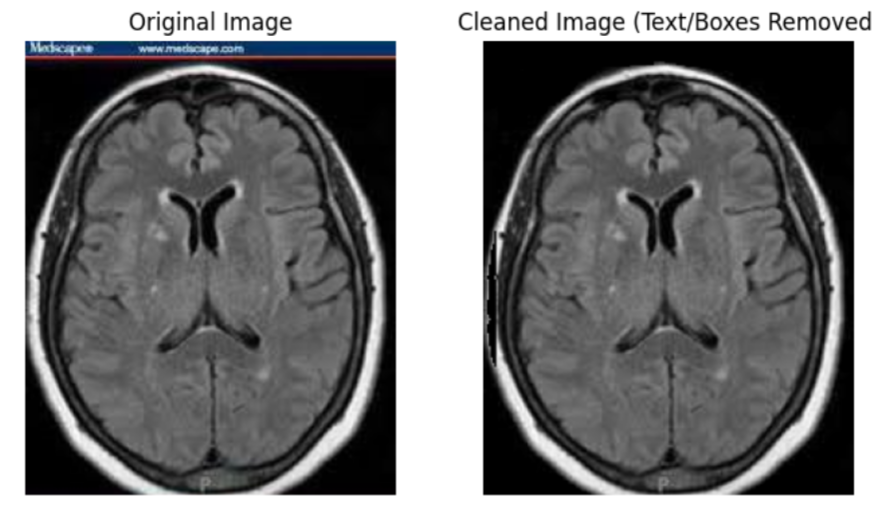
While this method had some limitations such as occasionally cropping the very edge of an MRI scan, it successfully removed the majority of problematic artifacts without affecting the brain region.
Lesson
The main lesson learnt was that rigorous exploratory data analysis (EDA) is a critical prerequisite for modeling in sensitive domains like medical imaging.
Thoroughly examining the dataset allowed us to identify quality issues that required special handling to ensure the final input was truly representative. Furthermore we realized that while cleaning is necessary, leaving minor artifacts can sometimes be beneficial as it forces the model to be more adaptive to real-world conditions where images are rarely perfect.
We also discovered that traditional cleaning methods were necessary because the bit depth of their JPEG dataset was incompatible with some specialized neuroimaging tools designed for other file formats.
Results
The models achieved the following results on their respective test sets:
Results - Vision Transformer (ViT)
This was the top-performing model, achieving a test accuracy of 96.25%.
It showed excellent precision and recall across all classes, with particularly high F1-scores for gliomas (0.98) and no tumor scans (0.98).
Results - Support Vector Machine (SVM)
Utilizing Histogram of Oriented Gradients (HOG) features, the SVM achieved a respectable test accuracy of 88.96%. It performed strongly in identifying gliomas and pituitary tumors but was less effective at distinguishing meningiomas.
Results - Convolutional Autoencoder (CAE)
This model reached an accuracy of 82.50%. It was highly reliable for “pituitary” (99.2% recall) and “notumor” (95.0% recall) classes but struggled significantly with meningiomas.
Results - Convolutional Neural Networks (CNN)
Two architectures were tested. Model 1 (Simple) reached 80.21% accuracy, while Model 2 (Complex) reached 80.63%. Like the other models, the CNNs were weakest at identifying meningiomas.
Results - General Insights
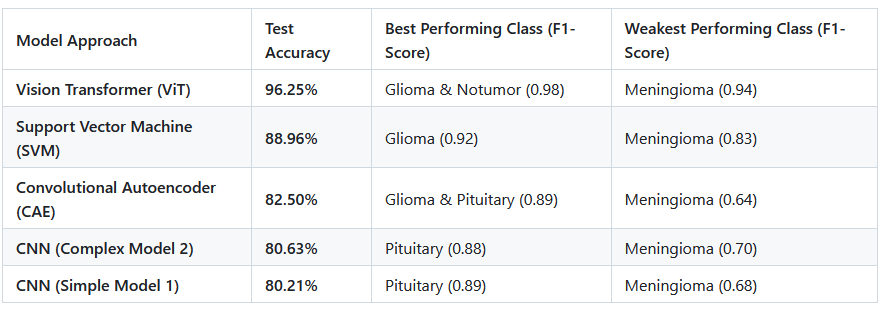
The Difficulty of Meningioma: Across almost all models, meningioma was the most challenging class to classify. For example, the CAE had a recall of only 51.7% for this tumor type, frequently confusing it with healthy tissue or pituitary tumors.
Feature Extraction vs. Automated Learning: The ViT’s success was attributed to its ability to automatically learn global spatial relationships and complex patterns directly from pixels, whereas the SVM relied on manual feature engineering (HOG), which may not have captured all critical diagnostic nuances.
Impact of Data Preparation: Because the initial dataset was small (120 images per class), the team used extensive data augmentation (e.g., elastic deformation, Gaussian noise, and zooming) to expand the training set to 2,317 images. This was crucial for preventing the deep learning models from overfitting.
Computational Trade-offs: While the ViT provided “state-of-the-art” accuracy, it was noted as being highly resource-intensive compared to more traditional models or simpler CNNs
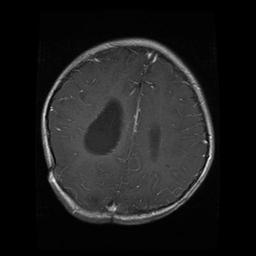 Brain Scan of Glioma Present
Brain Scan of Glioma Present
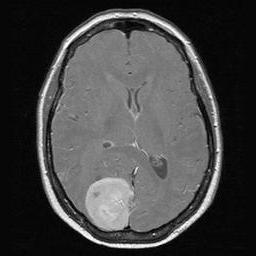 Brain Scan of Meningioma Present
Brain Scan of Meningioma Present
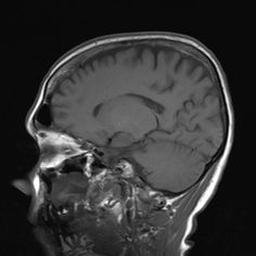 Brain Scan of No Tumor Present
Brain Scan of No Tumor Present
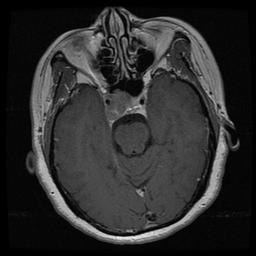 Brain Scan of Pituitary Present
Brain Scan of Pituitary Present
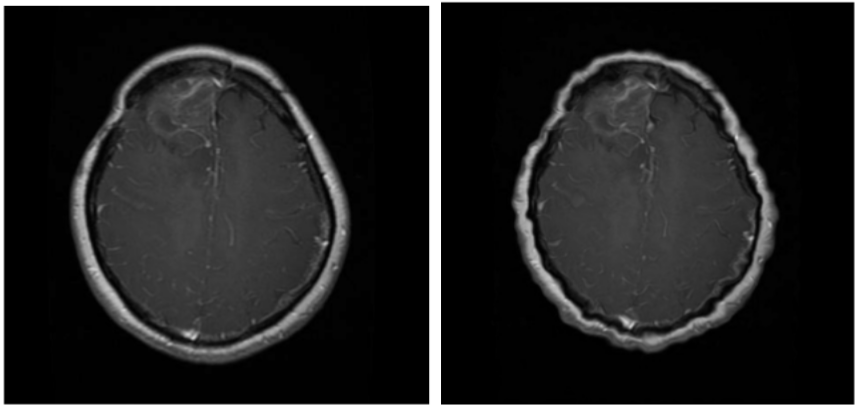 Elastic Deformation Applied - Before & After
Elastic Deformation Applied - Before & After
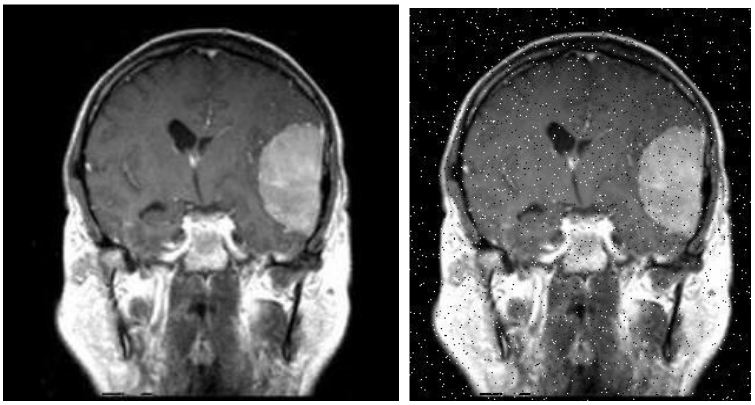 Salt & Pepper - Before & After
Salt & Pepper - Before & After
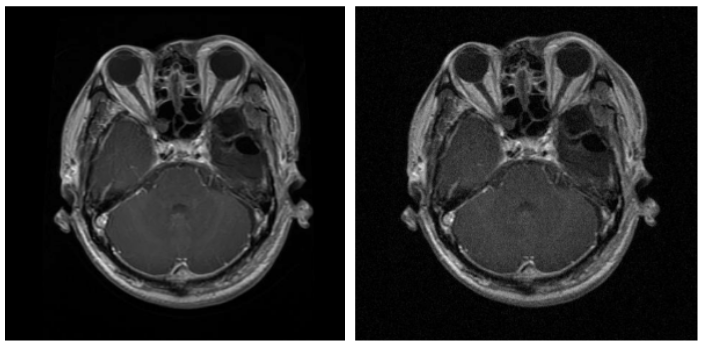 Gaussian Noise - Before & After
Gaussian Noise - Before & After
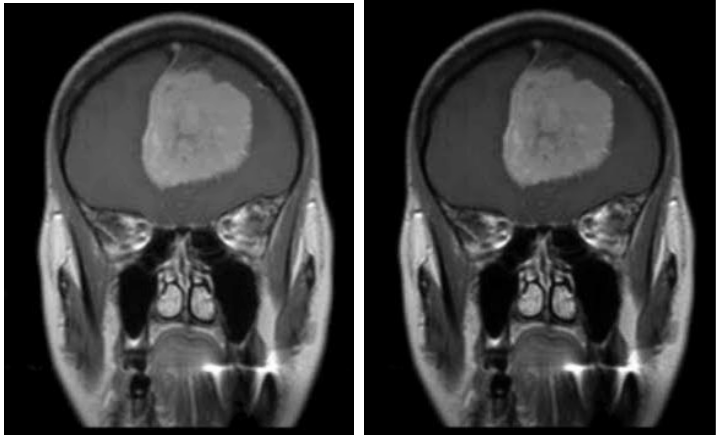 Gamma Correction - Before & After
Gamma Correction - Before & After
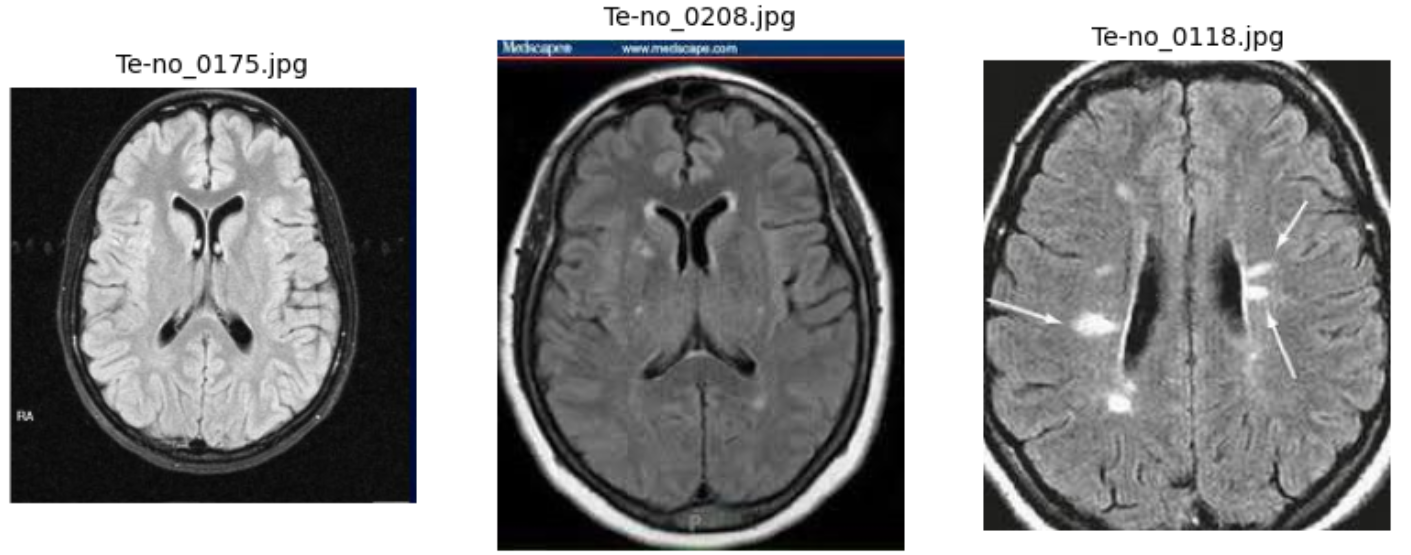 Example of Text Artifacts
Example of Text Artifacts
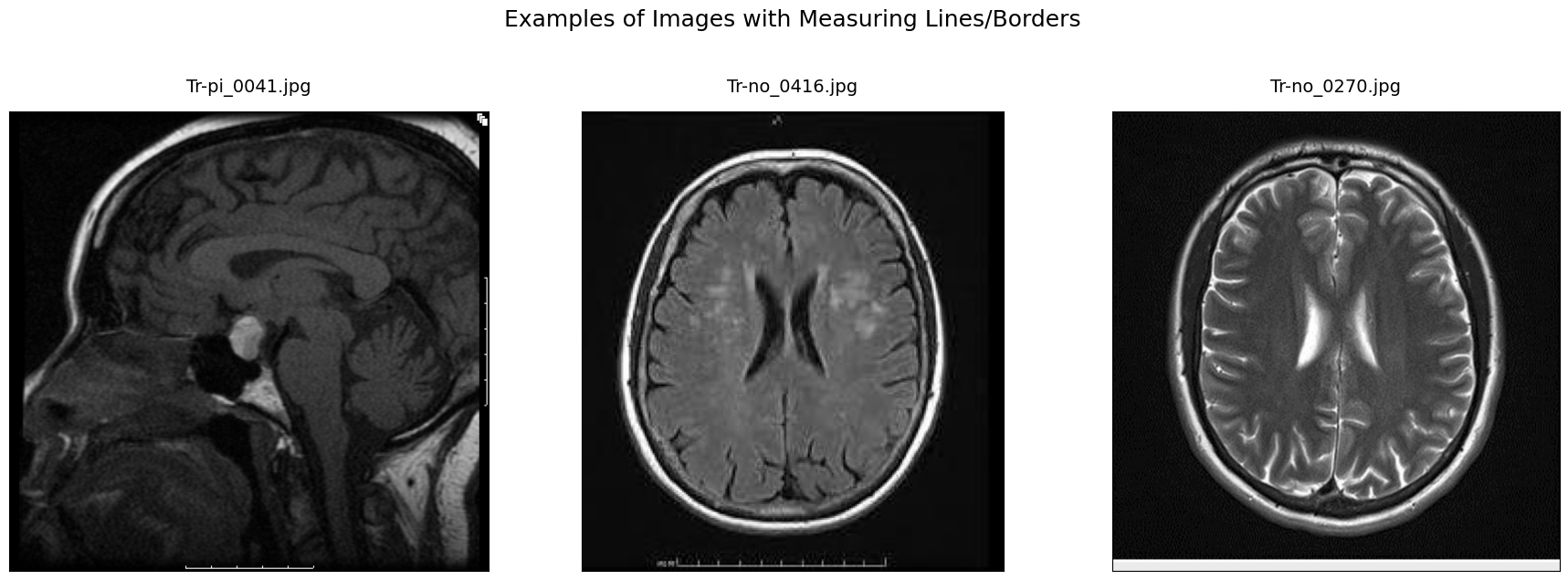 Example of Text Line Artifacts
Example of Text Line Artifacts
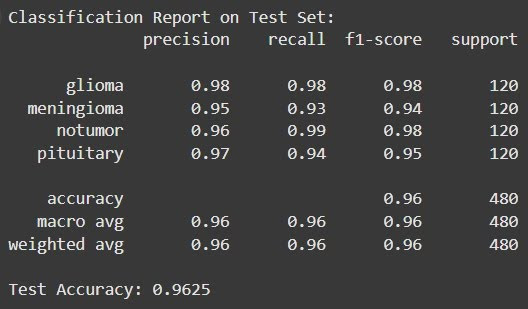 Test Evaluation Results
Test Evaluation Results
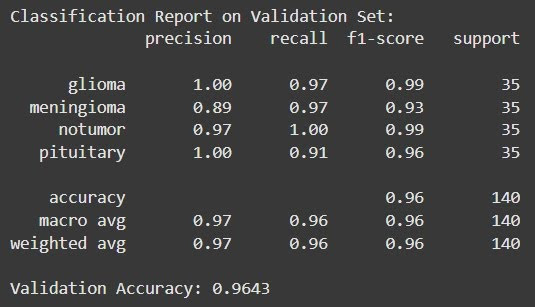 Validation Evaluation Results
Validation Evaluation Results
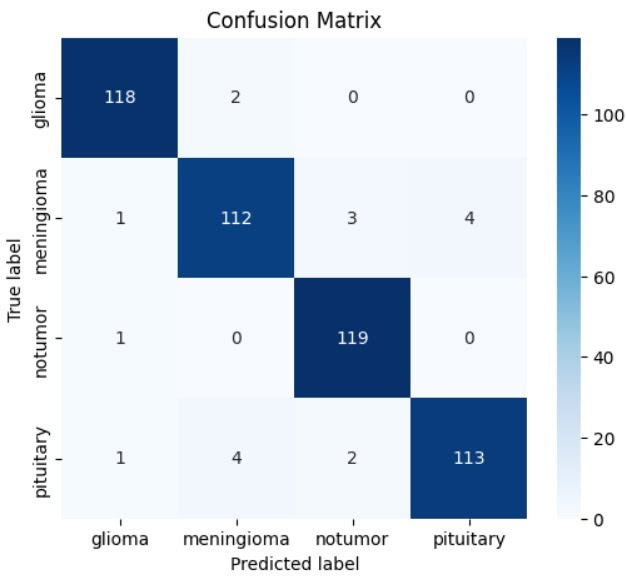 Confusion Matrix
Confusion Matrix
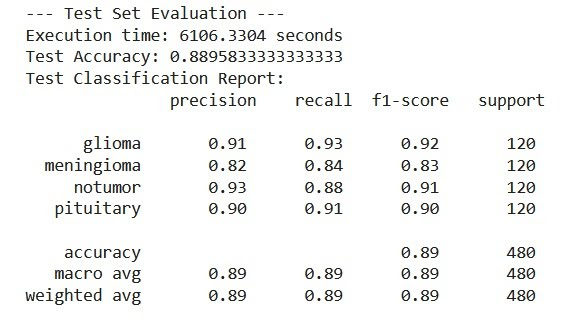 Test Evaluation Results
Test Evaluation Results
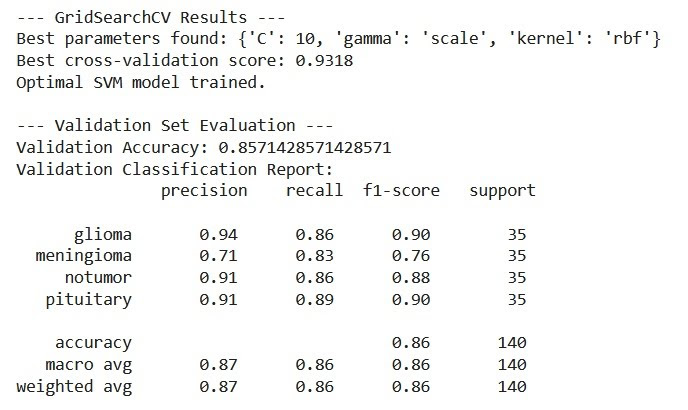 Validation Evaluation Results
Validation Evaluation Results
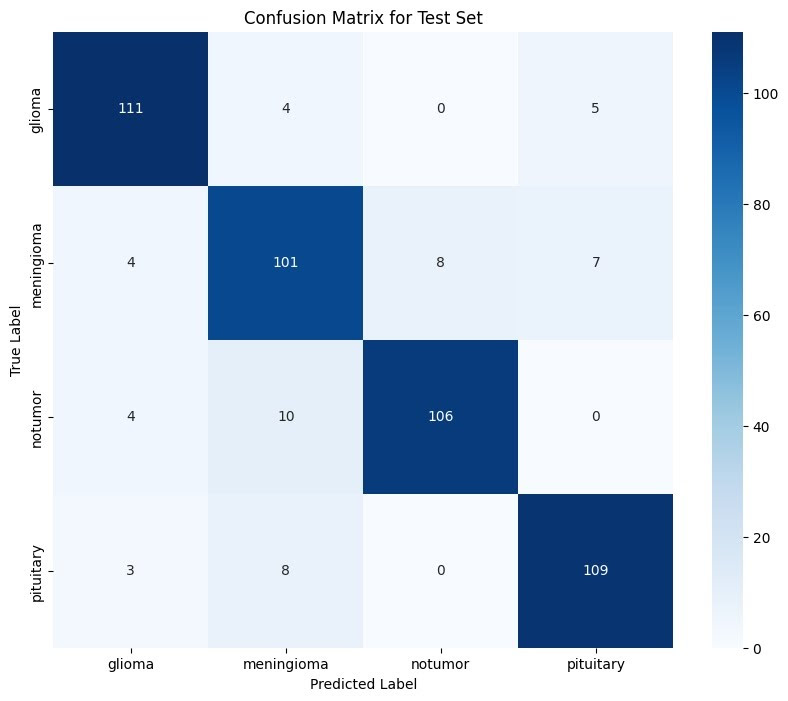 Confusion Matrix
Confusion Matrix
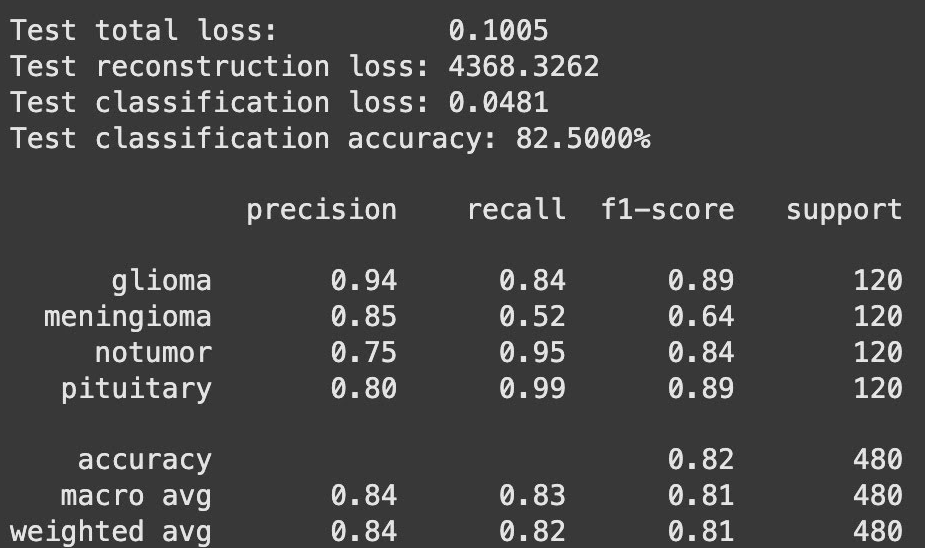 Classification Report
Classification Report
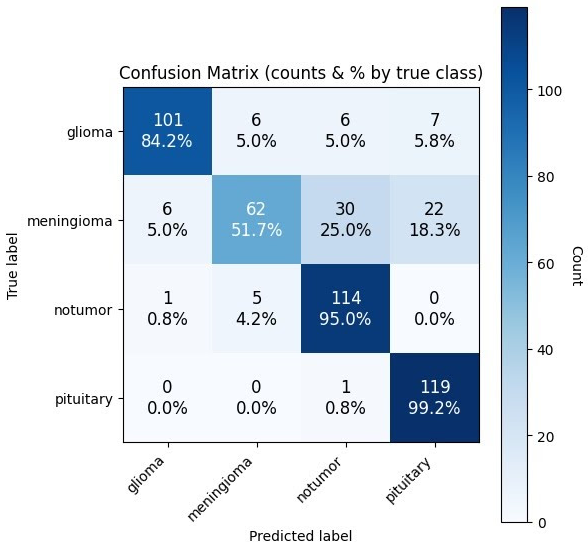 Confusion Matrix
Confusion Matrix
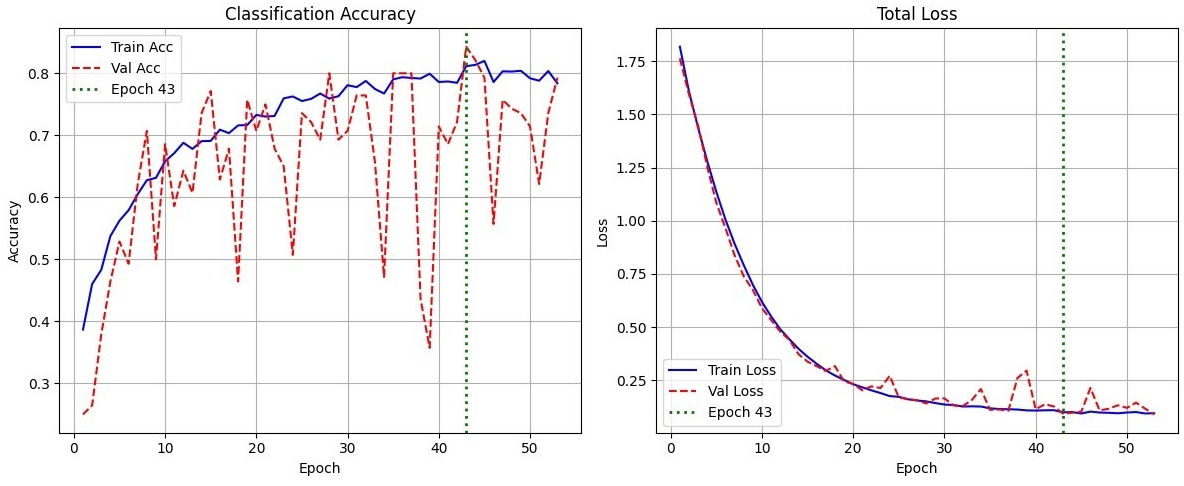 Training & Validation Accuracy
Training & Validation Accuracy
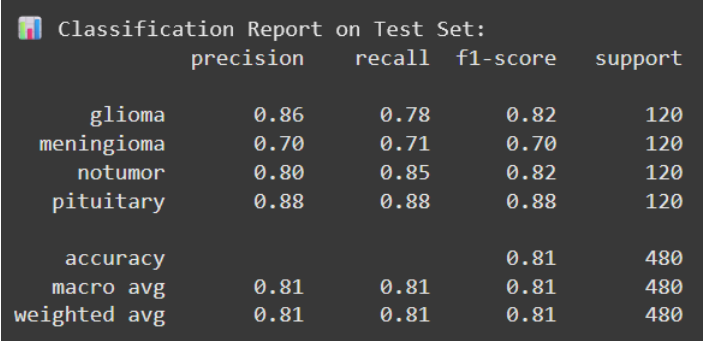 Model 1 - Test Classification Report
Model 1 - Test Classification Report
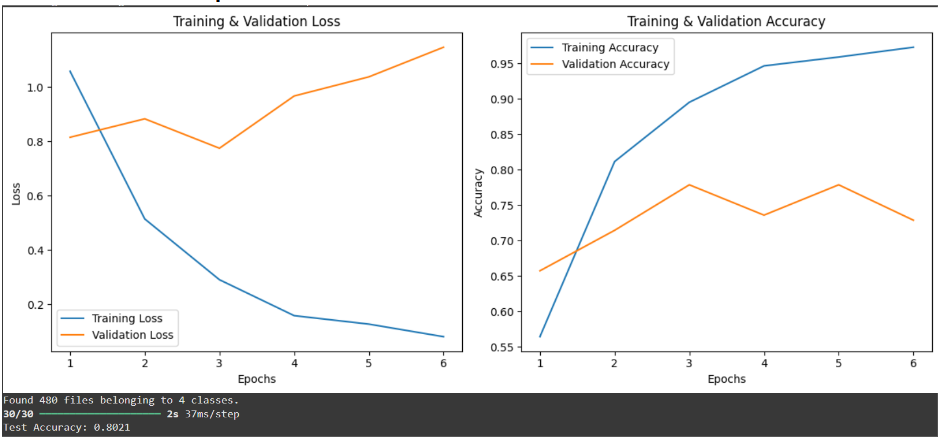 Model 1 - Training & Validation Loss/Accuracy
Model 1 - Training & Validation Loss/Accuracy
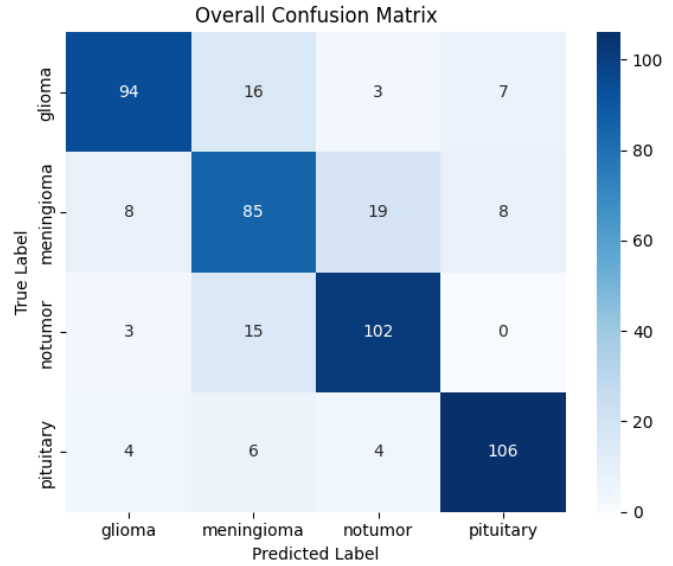 Model 1 - Confusion Matrix
Model 1 - Confusion Matrix
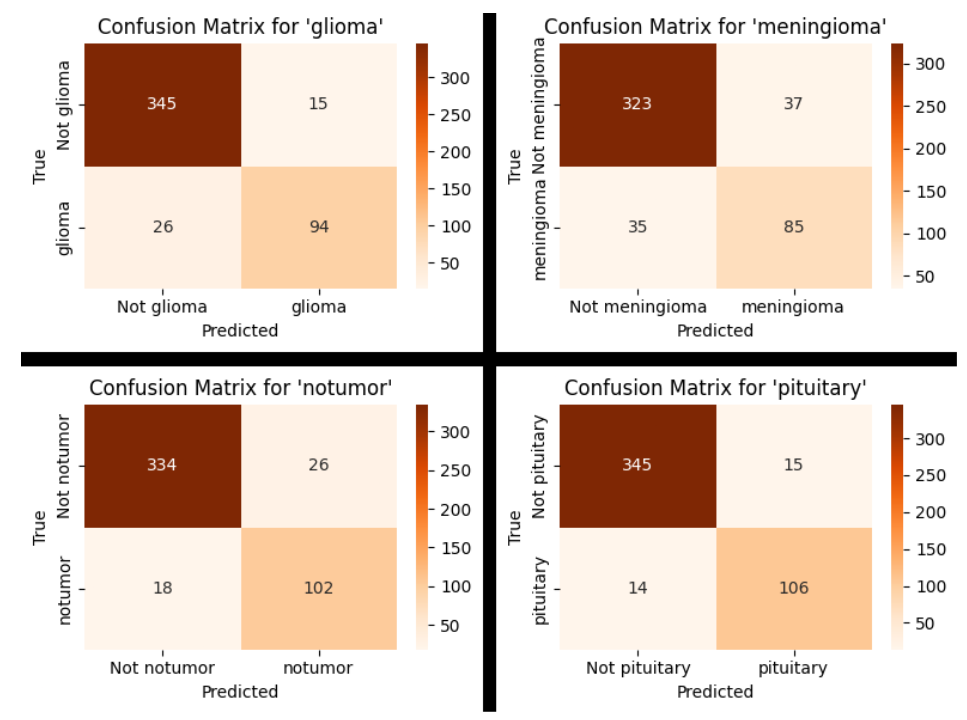 Model 1 - Confusion Matrix (By Tumour)
Model 1 - Confusion Matrix (By Tumour)
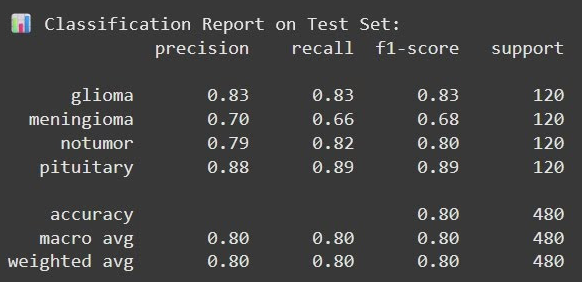 Model 2 - Test Classification Report
Model 2 - Test Classification Report
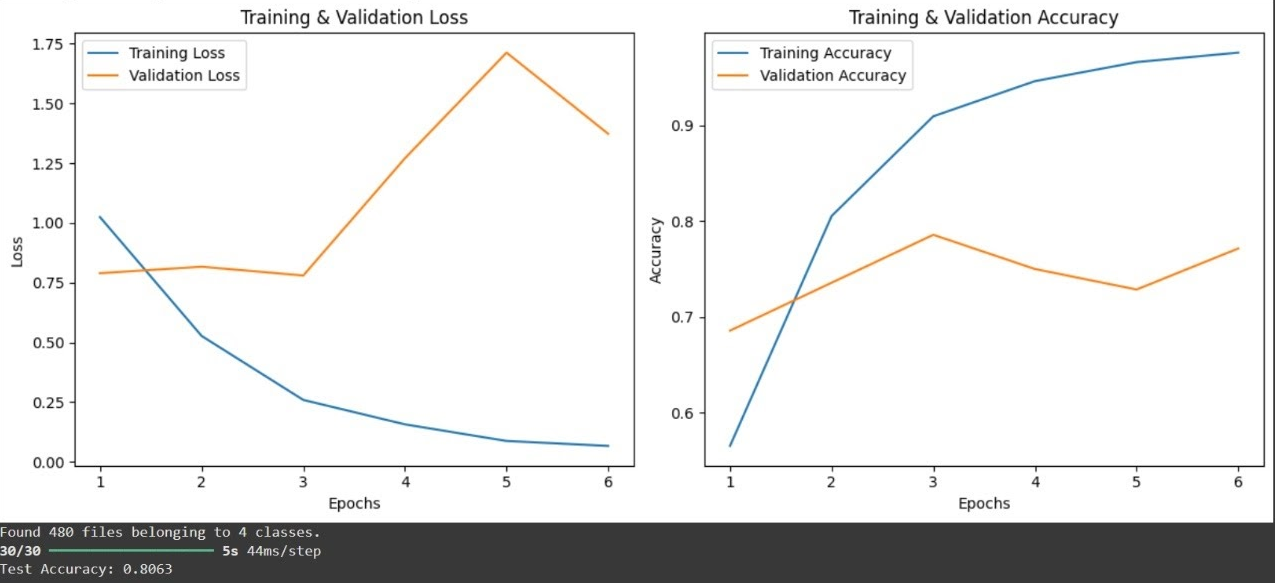 Model 2 - Training & Validation Loss/Accuracy
Model 2 - Training & Validation Loss/Accuracy
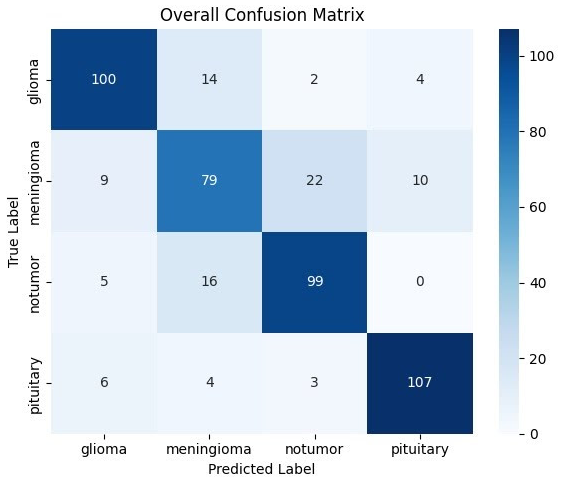 Model 2 - Confusion Matrix
Model 2 - Confusion Matrix
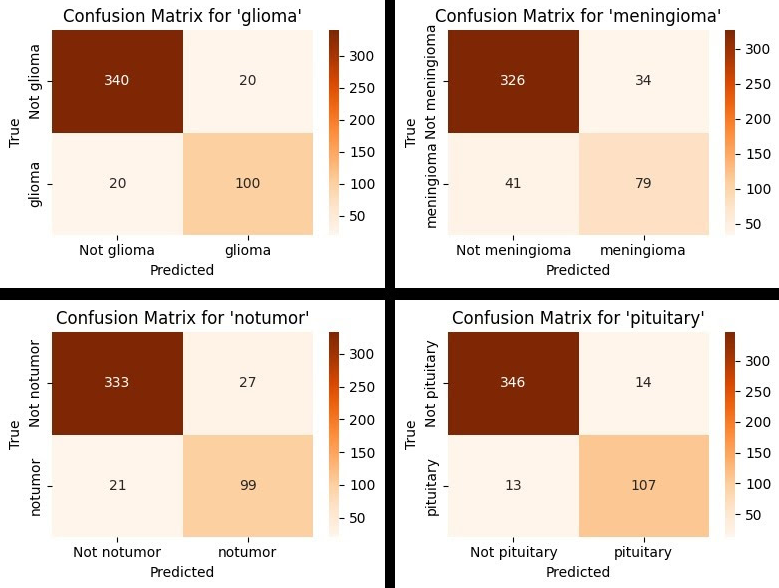 Model 2 - Confusion Matrix (By Tumour)
Model 2 - Confusion Matrix (By Tumour)